About Me

Welcome! I am a Zurich-based scientist focused on microbial systems biology. Here you will find the latest updates on my research, as well as my work on science accessibility.
Research
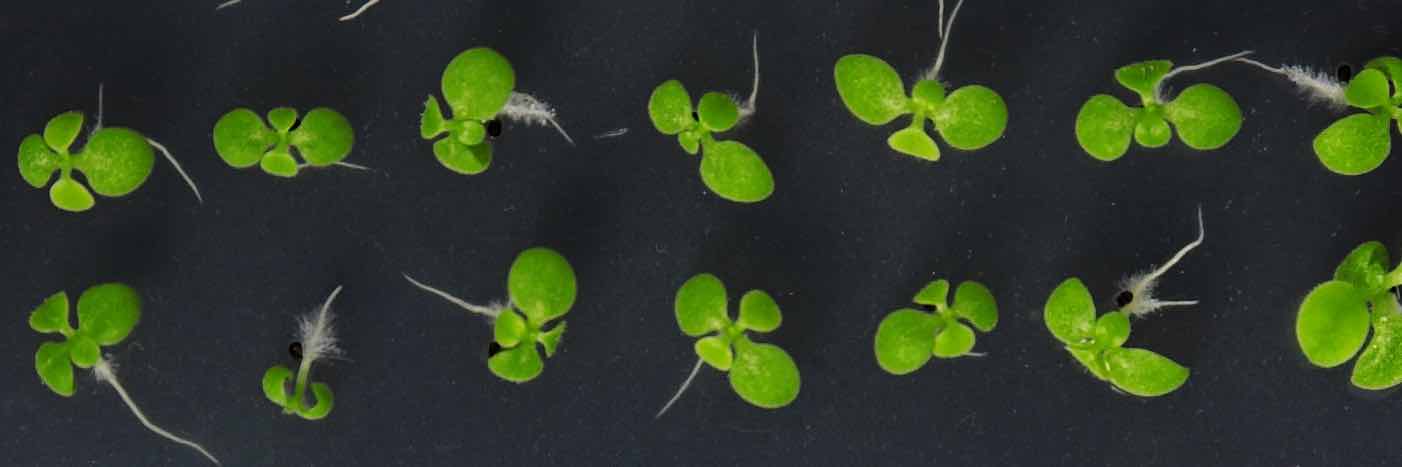
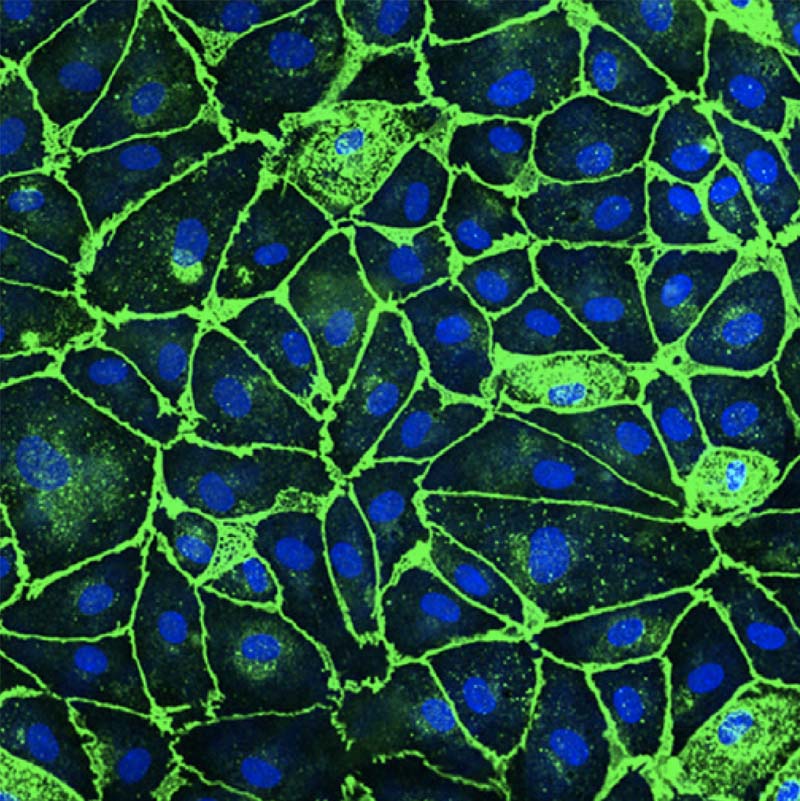
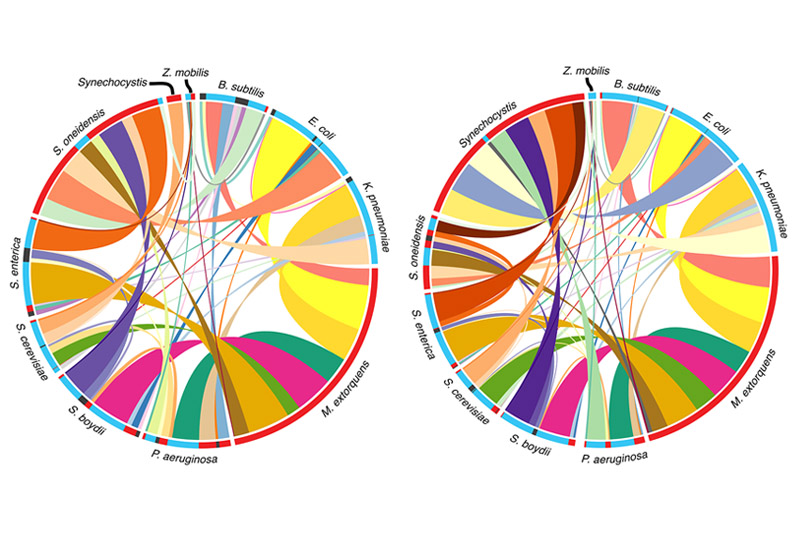
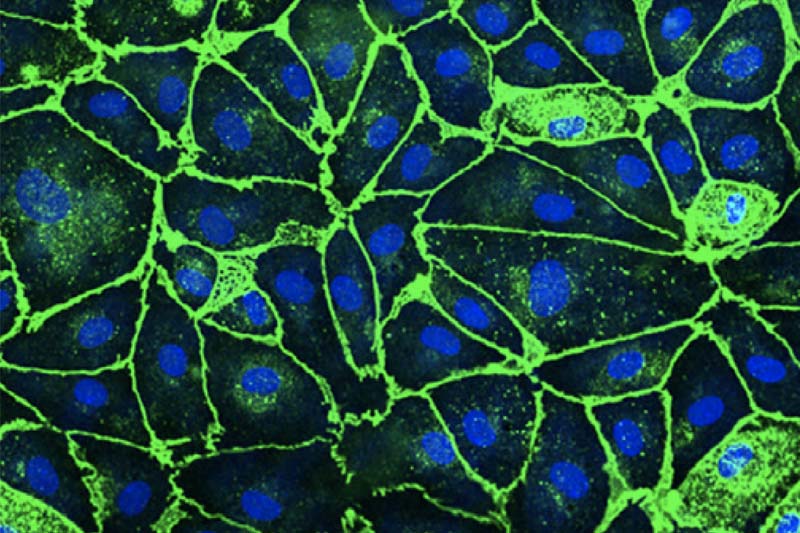
I am a JSMF postdoctoral fellow at ETH Zurich in the group of Julia Vorholt. My research is centered on microbial communities - the complex groups of tiny organisms that have enormous effects in all biomes: from the human gut to the ocean floor. Currently, I am focused on studying the microbial communities of plant leaves. Using a combination of experimental and modeling methods, I seek to better understand how these microbes interact with each other and with their environment, as well as how we can harness these interactions for applications in host and ecosystem health.
I completed my Ph.D. in bioinformatics at Boston University in the lab of Daniel Segrè, and was funded by a Howard Hughes Medical Institute Gilliam Fellowship and a National Academies Ford Fellowship.
Accessibility in science
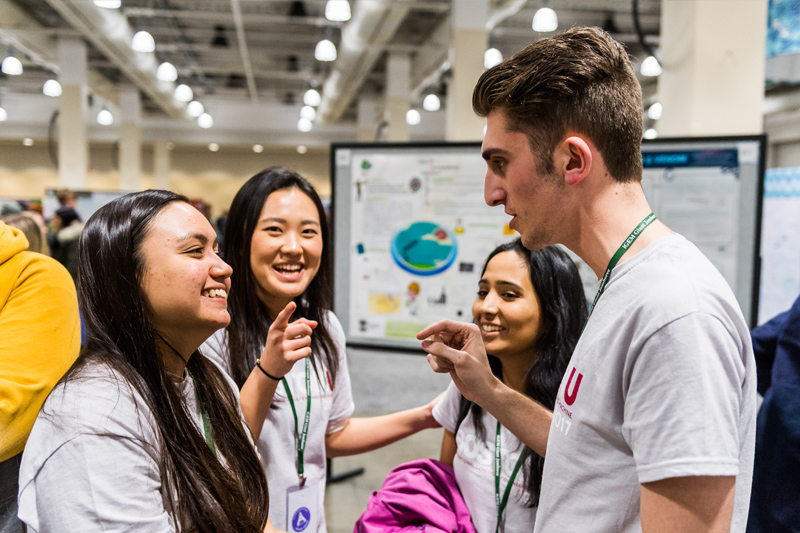
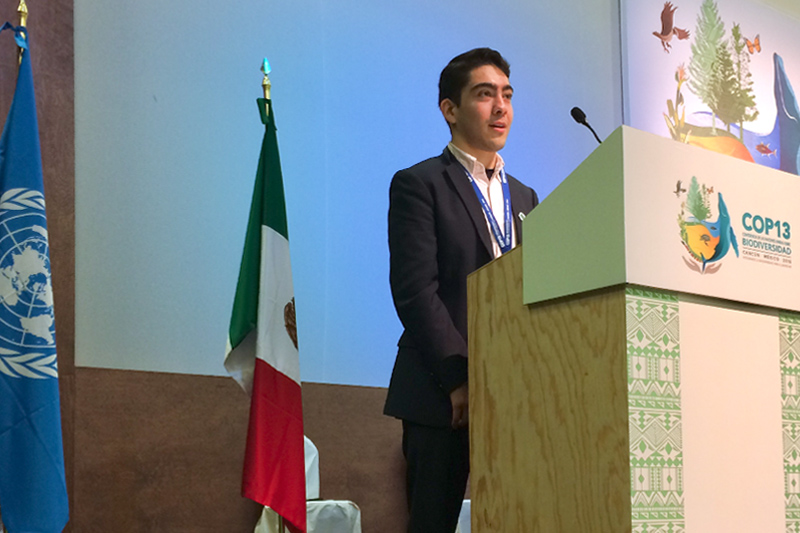
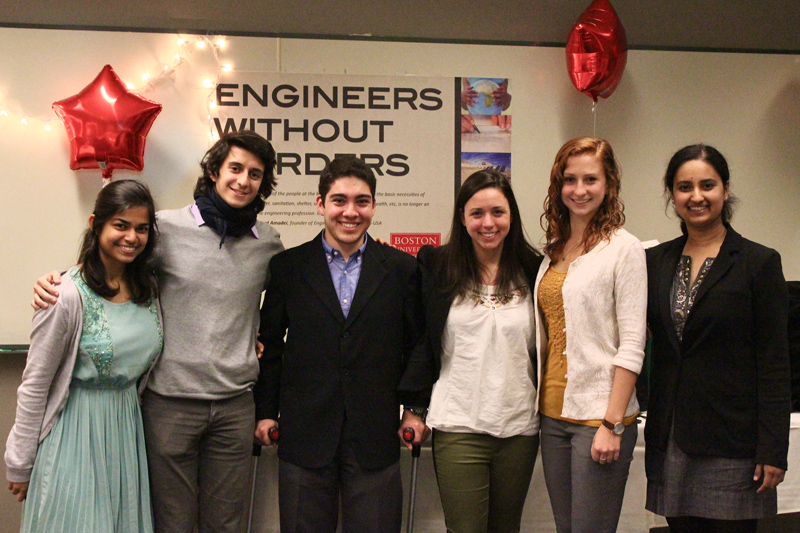
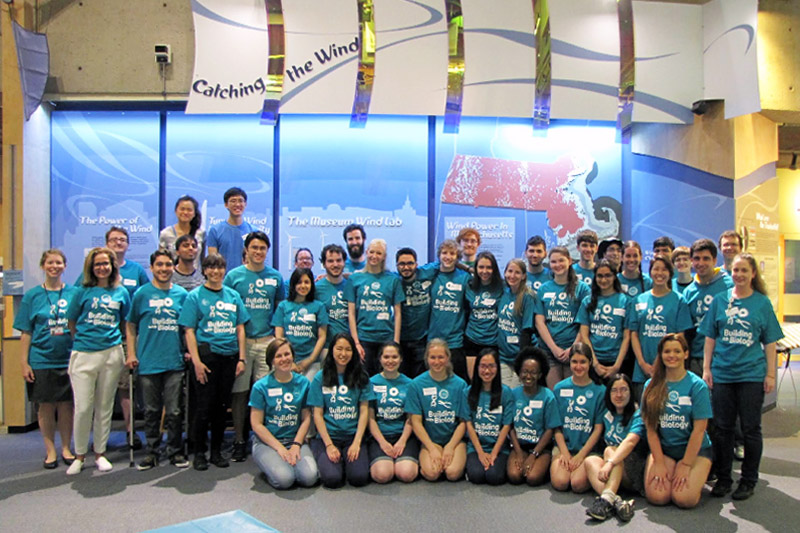
Science and technology impact all areas of our lives. As such, scientists have key roles to play in breaking down barriers of access to our own research, as well as to higher education. As part of this goal, I have worked on various programs aimed at creating and sustaining environments conducive to the success of students traditionally excluded from academia. I have also worked on a number of initiatives on the role of science in society, which have ranged from communicating microbiology to families in Boston and Zurich, to mentoring students in synthetic biology, to engaging with policymakers at the United Nations.
Read more about these projects here.
Graphic design is an excellent tool for bridging science and our own lives. Here, I've compiled a few art projects that explore this connection, as well as others I've made just for fun.